1. Introduction
Case-based Reasoning (CBR) solves new problems by retrieving previously solved problems and reusing the corresponding solutions. The specificity of the case-based approach of reasoning lies in its focus on the inseparability of reasoning from memory and from learning (Bichindaritz & Conlon, 1996) In CBR terminology, a case usually denotes a problem situation. A previously experienced situation, which has been captured and learned in a way that it can be reused in the solving of future problems, is referred to as a past case, previous case, stored case, or retained case. Correspondingly, a new case or unsolved case is the description of a new problem to be solved. Case-based reasoning is - in effect - a cyclic and integrated process of solving a problem, learning from this experience, solving a new problem, etc. (Aamodt & Plaza, 1994).
The common life cycle for solving a problem using CBR is mainly carried out in four-step process (Trendowicz & Jeffery, 2014):
Retrieve: One or more problems that are similar to the new problem are retrieves from the base of previously solved problems, and one attempts to modify them to fit the new problem parameters.
Reuse: The solutions of the selected previous problems are reused to solve the new one.
Revise: The solved new problem is then revised against the actual solution.
Retain: When successfully tested, it is added to the base of previous problems to be reused for solving future problems.
In particular, CBR has demonstrated to be an appropriate methodology for:
Working with unstructured domains data or difficult knowledge acquisition situation, for example, formal models or universally applicable guidelines do not well understand many diseases (Herrero, 2007), (Bichindaritz & Marling, 2006).
Help Desk systems used in the area of customer service to solve problems with products or services (Jenal, Gonzales, Alejo, & Ramos López, 2006).
Predictions of the possible success of a proposed solution when the information is stored considering the level of success of the solutions, the case-based reasoned can be able to predict proposed solution to the current problem. Clearly, the reasoner will have in not only those levels of success stored but also the differences between the Case or cases recovered and the current situation (Lozano & Fernández, 2008).
Automatic Acquisition of Subjective Knowledge because CBR systems exhibit an incremental knowledge acquisition, and knowledge can be abstracted by generalizing cases (Phuong, Hoang, Prasad, Hung, & Drake, 2001).
Medical diagnosis based on the similarity of the symptoms of the base of cases with the current case select the one that best suits to make a diagnosis (Jenal, Gonzales, Alejo, & Ramos López, 2006).
Among these tasks, medical diagnosis has been one of the most popular research subjects in both medical informatics and computer science communities. Medical diagnosis is the process aiming at identifying diseases based on findings (such as symptoms, lab reports, patient’s complaints, and other environmental factors) Adapting computer aided decision support systems to the diagnosis process requires a database-like medical knowledge base, and a problem-solving strategy applied to the knowledge base. A single problem solving strategy may be sufficient for simple diagnosis problems However, some difficulties may arise when the diagnosis problem becomes complex (Wang, Hsien-Tseng, & Tansel, 2013).
So, the CBR, is a reasoning process, which is medically accepted and also getting increasing attention from the medical domain. A number of benefits of applying CBR in the medical domain have already been identified (Bichindaritz & Marling, 2006), (Gierl, Bull, & Schmidt, 1998), (Montani, 2008). However, the medical applications offer several challenges for the CBR researchers and drive advances in research (Begum, Ahmed, Funk, Xiong, & Folke, 2011).
In order to adequately represent data and to avoid the inconveniences caused by its high dimensionality, we propose the use of variable selection and dimension reduction techniques in a preprocessing stage for CBR tasks, finally, we make a comparative study of multi-class classifiers to assess processed data performance.
The rest of this paper is structured as follows: Section II describes the proposed methodology, as well as the pattern recognition procedures used in this work. Section III presents the proposed experimental setup. Results and discussion are gathered in section IV. Finally, some concluding remarks and future works are drawn in Section V.
2. Material and Methods
This section outlines the proposed framework to assess the feasibility of using multi-class schemes within CBR approaches. Particularly, we resort to the adaptation of a pattern recognition stages into the CBR life cycle.
In the CBR scheme, the recovery is the most important stage, since in this phase the system finds the most similar cases to the current unknown case, simulating an efficient memory as a human expert would (Kolodner, 1983). By combining the CBR methodology with classifiers, a cost function would be used to find the nearby cases.
The next stage where we adapt classifiers would be in the adaptation stage, because we want to show the answer in terms of probabilities. With the classifier, we can find the membership degree of the new case in each of the classes, which would be helpful for medical staff.
To that end, we propose to carry out a comparative study of multi-class classifiers within preprocessing, recovery, and adaptation CBR stages. Fig. 1 depicts the proposed methodology to perform the comparison of multi-class classifiers.
Preprocessing
Variable selection: First, as preprocessing stage a variable selection procedure is employed. In this work, we use the so-called correlation based feature subset (CfsSubsetEval) algorithm, which evaluates the relevance of a subset of attributes by considering the individual predictive ability of each feature along with the degree of redundancy among them. Subsets of features that are highly correlated with the class while having low intercorrelation are preferred. And as search method the bestfirst algorithm, to reduce the number of parameters per instance of a dataset with a backtracking. It starts with the whole set of attributes and search backward to reduce the number of parameters per instance of a dataset.
Dimensionality Reduction: After performing variable selection and aiming to improve both visual inspection and classification performance, a dimensionality reduction stage is employed by using well known methods, namely Laplacian Eigenmaps (LE), that uses spectral techniques to perform dimensionality reduction. This technique relies on the basic assumption that the data lies in a low-dimensional manifold in a high-dimensional space (Belkin, 2003) and t-distributed stochastic neighbor embedding (t-SNE), it is a nonlinear dimensionality reduction technique that is particularly well-suited for embedding high-dimensional data into a space of two or three dimensions, which can then be visualized in a scatter plot.
Adaptation and recovery
Here, with the aim of accomplishing a multi-class case recovery, representative multi-class classifiers are considered. Due to their characteristics, we select the following classifiers: K Nearest Neighbor Classifier (K-NN) being a geometric-distance-based-approach, artificial neural networks (ANN) being a heuristic-search-based approach, support vector machines (SVM) being a model-based classifier, and Parzen's Classifier (PC) being a non-parametric density-based classifier.
3. Experimental Setup
A. Database
For evaluating the proposed methodology, we used two databases from UCI Machine Learning Repository. The first one, named Cardiotocograms, contains 2126 fetal cardiotocograms belonging to different classes. The dataset consists of measurements of fetal heart rate (FHR) and uterine contraction (UC) features on cardiotocograms classified by expert obstetricians. This data set consists of 21 attributes which include LB - FHR baseline (beats per minute), AC of accelerations per second, FM of fetal movements per second, UC of uterine contractions per second, DL of light decelerations per second, DS of severe decelerations per second, DP of prolonged decelerations per second, ASTV percentage of time with abnormal short term variability, MSTV mean value of short term variability, ALTV percentage of time with abnormal long term variability, MLTV mean value of long term variability, Width width of FHR histogram, Min minimum of FHR histogram, Max Maximum of FHR histogram, Nmax of histogram peaks, Nzeros of histogram zeros, Mode - histogram mode, Mean histogram mean, Median histogram median, Variance histogram variance, Tendency histogram tendency, CLASS FHR pattern class code (1 to 10) and NSP fetal state class code (Normal=1; Suspect=2; Pathologic=3).
The second database, named Cleveland, contains 303 instances. The "goal" field refers to the presence of heart disease in the patient. It is integer valued from 0 (no presence) to 4. Experiments with the Cleveland database have concentrated on simply attempting to distinguish presence (values 1,2,3,4) from absence (value 0). Consisting of 13 attributes which include age, sex, chest pain type (Typical angina=1; Atypical angina= 2; Non-anginal pain=3; Asymptomatic=4), resting blood pressure, cholesterol, fasting blood sugar (True=1; False=0), resting electrocardiographic results (Normal=1; Having ST-T wave abnormality (T wave inversions and/or ST elevation or depression of > 0.05 mV)=2; Showing probable or definite left ventricular hypertrophy by Estes' criteria=3), maximum heart rate, exercise induced angina (Yes=1; No=0), oldpeak, slope (Upsloping=1; Flat=2; Downsloping=3), number of vessels coloured, thal (Normal=3; Fixed defect=6; Reversible defect=7) and the classification values from 0 no presence to 4 types of heart diseases.
B. Parameter settings and procedures
As outcomes of the preprocessing stage, we obtain that Cardiotocograms database is reduced to 10 features, and Cleveland database to 7 features. Subsequently, as part of the same stage, by using dimensionality reduction techniques Cardiotocogram database is reduced to a 2-, 3-, 5-, 8-dimensional space. Likewise, Cleveland database is reduced to 2-, 3-, 5-dimensional space. As well, the whole subset of selected variables is considered for both databases.
For classification techniques, it should be stated out that a 20-fold cross-validation was performed to achieve unbiased results. Particularly, the following setup is established:
K-NN: Is a nonparametric supervised classification method based on distances. This instance-based classification technique needs a value for the number of neighbors (K), such parameter is optimized by means of a leave-one-out strategy, is K-fold cross validation taken to its logical extreme, with K equal to N, the number of data points in the set. That means that N separate times, the approximate function is trained on all the data except for one point and a prediction is made for that point.
ANN: The heuristic-based classification technique requires a number of units per hidden layer. In this work, a back-propagation trained feed-forward neural net is used with a single hidden layer. The number of units is computed from the data itself as the half of the instances divided by feature size plus the number of classes. The weight initialization consists of setting all weights to be zero, as well as the dataset is used as a tuning set.
SVM: This instance-based classification method takes advantage of the kernel trick to compute the most discriminative non-linear hyperplane between classes. Therefore, its performance heavily depends on the selection and tuning of the kernel type. For this work a Gaussian kernel is selected given its ability of generalization and its band-width parameter was fixed by the Silverman's rule (Sheather, 2004).
PC: This probabilistic-based classification method requires a smoothing parameter for the Gaussian distribution computation, which is optimized.
As a performance measure, it is used the standard mean classification error.
4. Results and Discussion
Achieved results for different number of dimensions as well as different classifiers are shown in Table I as the mean and standard deviation over the 20 folds runs. It can be seen how Cleveland dataset is a challenge task since performance is poor for all classifiers. It should be stated also that dimensionality reduction does not necessarily improves classification performance for both dimensionality reduction techniques. Nevertheless, by reducing dimensionality there is a gain in visual analysis of data as can be appreciated in Figure 1, particularly it can be seen how in 2D (Figures 2(a) and 2(c)) and 3D (Figures 2(b) and 2(d)) Cleveland data is highly overlapped which is consistent with achieved results. It should be noted that the error for SVM classifier is 0.397 ± 0.07, which is not far from the result obtained in (Bhatia, Praveen , & Pillai, 2008) and the Table 1, where the classification accuracy with 7 attributes is of 70.36%.
Table 1 Achieved classification performance over 20-fold cross validation for considered databases and dimensionality reduction techniques
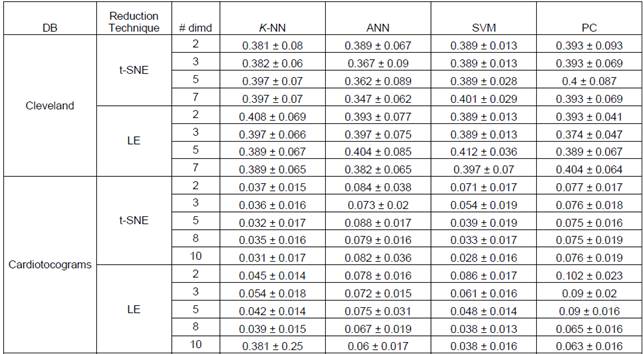
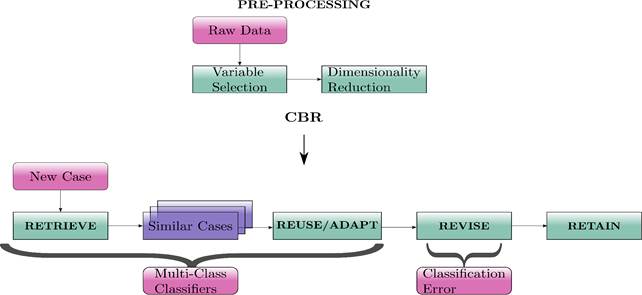
Fig. 1: Block diagram of proposed methodology. The aim of the comparative study is assessing the possibility of incorporating multi-class classifiers into CRB approaches design, as well as identifying the best classifier for this task
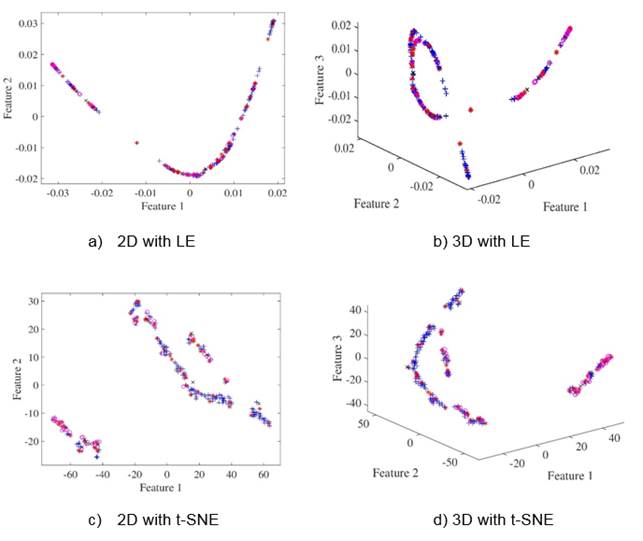
Fig. 2: Low-dimensional scatterplots for Cleveland database. Figures (a), (c) show the first two features from database. Figures (b), (d) show the first three features from database.
For cardiotocograms dataset classes separability is evident in lower dimensions, i.e. 2D and 3D, as depicted in Figures 3(a) to 3(d) leading to outstanding results as shown in Table I, however, as for Cleveland dataset, dimensionality reduction does not substantially improve classification performance on Cardiotocograms dataset even though it enhances data visualization. We can see that for the Cardiotocograms database the best result was using the SVM classifier the error is 0.028 ± 0.016, improving the results obtained in (Sundar, Chitradevi, & G. , 2012) where they achieved an average accuracy of 0.9328.
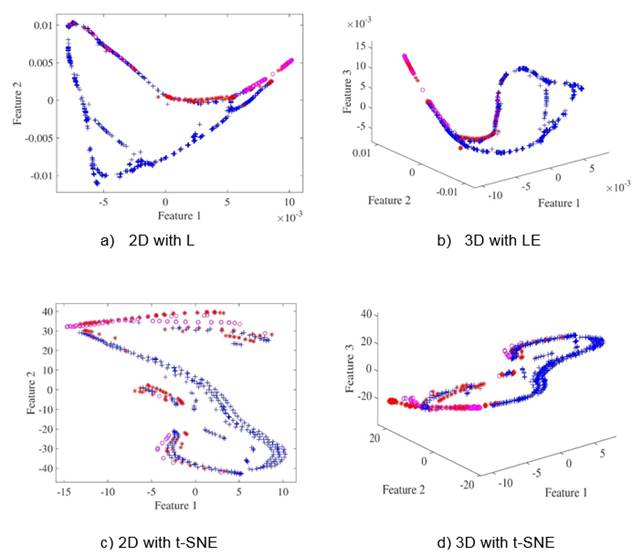
Figure 3 Low-dimensional scatterplots for Cardiotocograms database. Figures (a), (c) show the first two features from database. Figures (b), (d) show the first three features from database.
By performing a stability assessment, it could be seen from Figure 4 by the width of the error boxplots how SVM and K-NN classifiers achieves the best results for considered Cardiotocogram and Cleveland databases. Moreover, it should be noted how SVM classification results are the most stable of the considered classification techniques.
5. Conclusions and future work
This work presents a feasibility evaluation of the use of techniques from the field of pattern recognition into CBR frameworks, so that conventional CBR can be extended to multi-class scenarios. The above, with the aim of facilitating decision making in the diagnostic support, especially in situations where subcategories may exist and even emerging categories according to the condition of the patients.
Experimentally we prove that the SVM classifier is a good candidate for integration with the CBR approach to create a generic system to assist physicians in the diagnosis of patients and is capable of working with databases multiclass associating probabilities each class, responding to one of the challenges of (Bichindaritz & Marling, 2006), (Kwiatkowska & Atkins, 2004).
As a future work, we will explore the possibility to design a case recovery stage for CBR able to deal with multi-class cases while providing users with class membership (probabilities to belong) estimates for a new case, improving usability with respect to conventional approaches and, in addition, providing more meaningful information to the expert at the review stage in order to provide a more accurate diagnosis.
To improve the performance of the classifiers, we will study new techniques of variable selection and reduction of dimensions, as well as techniques of data balancing for databases that require it as is the case of Cleveland, due to problems such as class overlapping or the lack of data representative of input data.